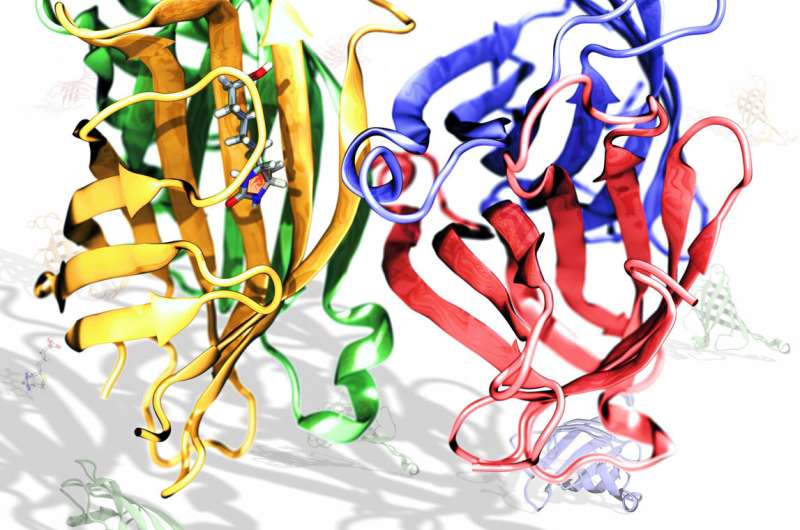
Natural frequencies of the α-actinin dimer. G) Image showing the torsional movement in mode 11. Arrows point to location of the three hinges. F) Image captures movement characteristic of the three-hinge bending modes (9, 10, and 12). Arrow points to location of the single-hinge. E) Image captures movement characteristic of the single-hinge bending modes (7 and 8). In the α-actinin monomer normal mode analysis only mode 11 showed torsional movement. D) Vector field representation of movement in the torsional mode. Mode 9, mode 10, and mode 12 showed three-hinge bending movement. C) Vector field representation of movements during three-hinge bending modes. Mode 7 and mode 8 exhibited this type of bending motion. Larger vectors represent larger movements. Vector field representations show the direction and magnitude of movement of each residue in the molecule. B) Vector field representation of movements during one-hinge bending normal modes. Single-hinge bending modes are shown in blue colors, three-hinge bending modes in green colors, and the torsional mode is shown in pink. Natural movements consisted mainly of movement in the regions near the termini. Normal mode analysis of the α-actinin monomer revealed bending and torsional natural frequencies.Ī) RMSD of individual residues for six lowest vibrational modes. In dimer conformation R1 is interacting with R4 and R2 is interacting with R3. R1 is colored in red, R2 in yellow, R3 in green, and R4 in blue. Each of the four spectrin repeats are colored according to conformation. B) VMD generated image of the α-actinin dimer rod domain. The monomers are arranged in an anti-parallel manner. tilt: cosine of the rotation orthogonal to a given axis.A) α-actinin is a dimer with three major domains: an actin binding domain, a calmodulin homology domain, and a central rod domain.spinAngle: angle of rotation around a given axis.orientationProj: cosine of the angle of rotation from reference coordinates.orientationAngle: angle of rotation from reference coordinates.orientation: orientation from reference coordinates.inertiaZ: total moment of inertia of a group of atoms around a chosen axis.inertia: total moment of inertia of a group of atoms.gyration: radius of gyration of a group of atoms. 
eigenvector: projection of the atomic coordinates on a vector.rmsd: root mean square displacement (RMSD) from reference positions.selfCoordNum: coordination number between atoms within a group.coordNum: coordination number between two groups.dihedral: torsional angle between four groups.dipoleAngle: angle between two groups and dipole of a third group.cartesian: vector of atomic Cartesian coordinates.distancePairs: set of pairwise distances between two groups.distanceInv: mean distance between two groups of atoms.distanceDir: distance unit vector between two groups.distanceVec: distance vector between two groups.distanceXY: modulus of the projection of a distance vector on a plane.distanceZ: projection of a distance vector on an axis.distance: center-of-mass distance between two groups.Collective variable components (basis functions).Performance a Colvars calculation based on group size.Treatment of periodic boundary conditions.Selecting atoms for colvars: defining atom groups.Statistical analysis of collective variables.General options for a collective variable.Defining collective variables and their properties.Configuration syntax for the Colvars module.General parameters and input/output files.Collective Variables Interface (Colvars).Coloring Trick - Override a Coloring Category.Creating a set of black-and-white color definitions.Using the Python interpreter within VMD.Implicit Ligand Sampling ( volmap ils command).Turn off highlighting / Change highlight style.Selecting residues by clicking on the 3-D structure.Selecting residues from the Sequence window listing.Display Menu and Display Settings Window.Using the Joystick in the Graphics Window.Using the Spaceball in the Graphics Window.Tracking Script Command Versions of the GUI Actions.Parallel Computing on Clusters and Supercomputers.Basic Hardware and Software Requirements. 
VMD development is supported by the National Institutes of Health 
Use the scripting interfaces for analysis and to This guide documents the user interfacesĭisplaying and grapically manipulating molecules, and describes how to How to run and use the molecular visualization and analysis University of Illinois at Urbana-Champaign NIH Biomedical Research Center for Macromolecular Modeling and Bioinformaticsīeckman Institute for Advanced Science and Technology
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
